In a paper in the journal Science, a team led by Professor Michael Yartsev’s lab identified the part of the brain in Egyptian fruit bats that controls vocalizations and found that it contains very similar neural wiring to the part of the human brain that controls speech.
research
Microfluidics: Biology’s Liquid Revolution
Professor Aaron Streets was featured in this overview on the potential of microfluidics in The Scientist magazine.
Rubinsky’s coral preservation work featured on PBS News
Professor Emeritus Boris Rubinsky’s isochoric vitrification method of preserving coral samples in suspended animation is part of recent emergency efforts to save dying coral reefs. The method is being used by the Smithsonian’s National Zoo and Conservation Biology Institute.
The booming business of discovering your biological age
Professor Irina Conboy and former student Alina Su have founded a new company, Generation Lab, offering an at-home molecular aging test that analyzes a person’s biological age by assessing “biological noise” in their system. The test evaluates an individual’s risk for top health conditions and the pace of aging across 19 systems in the body, which can help physicians see where interventions may be most needed and effective.
Researchers make advances toward more effective IBD therapies
Researchers in Professor Phillip Messersmith’s lab have demonstrated that treatment with DPCA, an enzyme inhibitor molecule shown to trigger regeneration in mammals, can protect against and repair colon damage in a mouse model of colitis. This work suggests that short-term use of this small molecule drug could someday provide a restorative therapy for patients with IBD — and a path to remission.
Herr Lab receives grant to study marine symbiosis in a warming world
The Herr Lab has been awarded a 3-year ‘Symbiosis’ grant from the Gordon & Betty Moore Foundation, geared towards designing and disseminating microfluidic tools to power new understanding of marine symbiotic systems – like coral reefs – adversely impacted by rising sea temperatures and other climate-associated stresses. Herr’s lab welcomes two new postdoctoral scholars, Drs. Fangchen Liu and Cyril Deroy, and is collaborating with experts in coral systems from the Carnegie Institution for Science (Prof. Phillip Cleves) and the University of Miami (Prof. Nikki Traylor-Knowles).
Undergrad Tau presents at ABRCMS
Undergraduate researcher Cyrus Tau was selected to present, and won the Best Talk award, in the Biochemistry/Molecular Biology oral presentation section at this year’s Annual Biomedical Research Conference for Minoritized Scientists – ABRCMS 2023.
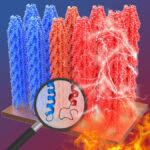
Researchers demonstrate heat-induced pyroelectricity in viruses
Researchers in Professor Seung-Wuk Lee’s lab discovered for the first time “heat-induced electrical potential generation on a virus,” a phenomenon known as pyroelectricity. This work may shed light on how biomaterials — cells, tissues and proteins — generate electricity at a molecular level as well as lead to the development of biomaterials with novel medical, pharmaceutical, environmental and energy applications.
How scientists aim to extend human lifespans
Professor Irina and Mike Conboy were guests on the Economist “Babbage” science and technology podcast dicussing progress in research on aging.